Characterization, Evolutionary Insights, and Stress-Responsive Expression of the Phosphoenolpyruvate Carboxylase (PEPC) Gene Family in Salix Matsudana

Article Information
Abstract
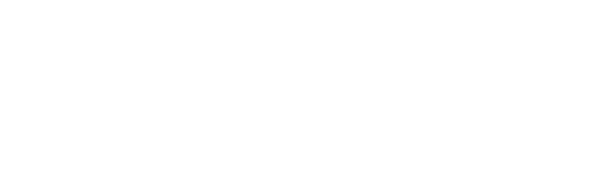
Phosphoenolpyruvate carboxylase (PEPC) is a vital enzyme in plant carbon metabolism, catalyzing the conversion of phosphoenolpyruvate (PEP) to oxaloacetate (OAA), a key intermediate in numerous biosynthetic pathways. PEPC family members are usually classified into PTPC and BTPC subfamilies, and PTPC subfamily is characterized by a conserved N-terminal serine phosphorylation site, while BTPC lacks this site. In this study, we identified 10 Salix matsudana PEPC genes (SmPEPCs), which were classified into two subfamilies, PTPC and BTPC, based on phylogenetic tree. Our findings highlighted significant gene duplication events that contributed to the expansion of the SmPEPC gene family, shedding light on its evolutionary development in willow. Comparative synteny analysis with Arabidopsis thaliana, Oryza sativa, Populus trichocarpa, and S. purpurea revealed both conserved and diverged patterns of PEPC organization across species. Interaction network analysis, along with Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis, indicated that SmPEPCs are involved in crucial metabolic pathways related to carbon flux. Expression profiling under salt or submergence stress revealed distinct expression patterns of SmPEPCs. Specifically, SmPEPC9 showed predominant upregulation under salt stress, while SmPEPC6 was mainly upregulated in submergence-tolerant varieties under submergence stress. These results suggested that SmPEPCs potentially modulate carbon flux and organic acid metabolism to mitigate stress effects, playing a central role in stress adaptation mechanisms.
Graphical Abstract
Keywords
Data Availability Statement
Funding
Conflicts of Interest
AI Use Statement
Ethical Approval and Consent to Participate
References
- O'Leary, B., Park, J., & Plaxton, W. C. (2011). The remarkable diversity of plant PEPC (phosphoenolpyruvate carboxylase): recent insights into the physiological functions and post-translational controls of non-photosynthetic PEPCs. Biochemical Journal, 436(1), 15-34.
[CrossRef] [Google Scholar] - Lucius, S., & Hagemann, M. (2024). The primary carbon metabolism in cyanobacteria and its regulation. Frontiers in Plant Science, 15, 1417680.
[CrossRef] [Google Scholar] - Westers, H., Dorenbos, R., Van Dijl, J. M., Kabel, J., Flanagan, T., Devine, K. M., ... & Quax, W. J. (2003). Genome engineering reveals large dispensable regions in Bacillus subtilis. Molecular Biology and Evolution, 20(12), 2076-2090.
[CrossRef] [Google Scholar] - Monreal, J. A., McLoughlin, F., Echevarría, C., García-Mauriño, S., & Testerink, C. (2010). Phosphoenolpyruvate carboxylase from C4 leaves is selectively targeted for inhibition by anionic phospholipids. Plant Physiology, 152(2), 634-638.
[CrossRef] [Google Scholar] - Cousins, A. B., Baroli, I., Badger, M. R., Ivakov, A., Lea, P. J., Leegood, R. C., & Von Caemmerer, S. (2007). The role of phosphoenolpyruvate carboxylase during C4 photosynthetic isotope exchange and stomatal conductance. Plant Physiology, 145(3), 1006-1017.
[CrossRef] [Google Scholar] - Ludwig, M. (2016). The roles of organic acids in C4 photosynthesis. Frontiers in Plant Science, 7, 647.
[CrossRef] [Google Scholar] - Nguyen, T. B. A., Lefoulon, C., Nguyen, T. H., Blatt, M. R., & Carroll, W. (2023). Engineering stomata for enhanced carbon capture and water-use efficiency. Trends in Plant Science, 28(11), 1290-1309.
[CrossRef] [Google Scholar] - Tan, B., & Chen, S. (2023). Defining mechanisms of C3 to CAM photosynthesis transition toward enhancing crop stress resilience. International Journal of Molecular Sciences, 24(17), 13072.
[CrossRef] [Google Scholar] - Li, C., Wang, J., Lan, H., & Yu, Q. (2024). Enhanced drought tolerance and photosynthetic efficiency in Arabidopsis by overexpressing phosphoenolpyruvate carboxylase from a single-cell C4 halophyte Suaeda aralocaspica. Frontiers in Plant Science, 15, 1443691.
[CrossRef] [Google Scholar] - Caburatan, L., & Park, J. (2021). Differential expression, tissue-specific distribution, and posttranslational controls of phosphoenolpyruvate carboxylase. Plants, 10(9), 1887.
[CrossRef] [Google Scholar] - Hu, R., Yu, H., Deng, J., Chen, S., Yang, R., Xie, H., ... & Yu, Y. (2025). Phosphoenolpyruvate and related metabolic pathways contribute to the regulation of plant growth and development. International Journal of Molecular Sciences, 26(1), 391.
[CrossRef] [Google Scholar] - Cao, J., Cheng, G., Wang, L., Maimaitijiang, T., & Lan, H. (2021). Genome-wide identification and analysis of the phosphoenolpyruvate carboxylase gene family in Suaeda aralocaspica, an annual halophyte with single-cellular C4 anatomy. Frontiers in Plant Science, 12, 665279.
[CrossRef] [Google Scholar] - Xu, R., Liu, H., Liu, C., Xia, M., Feng, D., Zhu, Y., ... & Chen, Z. (2025). Genome-wide identification and expression analysis of the PEPC gene family in Zanthoxylum armatum reveals potential roles in environmental adaptation. Biology, 14(11), 1605.
[CrossRef] [Google Scholar] - Lepiniec, L., Vidal, J., Chollet, R., Gadal, P., & Crétin, C. (1994). Phosphoenolpyruvate carboxylase: structure, regulation and evolution. Plant Science, 99(2), 111-124.
[CrossRef] [Google Scholar] - Wang, N., Zhong, X., Cong, Y., Wang, T., Yang, S., Li, Y., & Gai, J. (2016). Genome-wide analysis of phospho enol pyruvate carboxylase gene family and their response to abiotic stresses in soybean. Scientific reports, 6(1), 38448.
[CrossRef] [Google Scholar] - Feria, A. B., Bosch, N., Sánchez, A., Nieto-Ingelmo, A. I., de la Osa, C., Echevarría, C., ... & Monreal, J. A. (2016). Phosphoenolpyruvate carboxylase (PEPC) and PEPC-kinase (PEPC-k) isoenzymes in Arabidopsis thaliana: role in control and abiotic stress conditions. Planta, 244(4), 901-913.
[CrossRef] [Google Scholar] - Shaheen, I., Waseem, M., Basharat, S., Pingwu, L., & Qayyum, A. (2025). Bioinformatic identification of phosphoenolpyruvate carboxylase (PEPC) gene family in Brassica napus. Genetic Resources and Crop Evolution, 72(5), 5295-5309.
[CrossRef] [Google Scholar] - Zhao, Y., Guo, A., Wang, Y., & Hua, J. (2019). Evolution of PEPC gene family in Gossypium reveals functional diversification and GhPEPC genes responding to abiotic stresses. Gene, 698, 61-71.
[CrossRef] [Google Scholar] - Punyasu, N., Kalapanulak, S., & Saithong, T. (2023). CO2 recycling by phospho enol pyruvate carboxylase enables cassava leaf metabolism to tolerate low water availability. Frontiers in Plant Science, 14, 1159247.
[CrossRef] [Google Scholar] - Zhang, Z., Zhang, A., Zhang, Y., Zhao, J., Wang, Y., Zhang, L., & Zhang, S. (2024). Ectopic expression of HaPEPC1 from the desert shrub Haloxylon ammodendron confers drought stress tolerance in Arabidopsis thaliana. Plant Physiology and Biochemistry, 208, 108536.
[CrossRef] [Google Scholar] - Liu, D., Hu, R., Zhang, J., Guo, H. B., Cheng, H., Li, L., ... & Yang, X. (2021). Overexpression of an agave phosphoenolpyruvate carboxylase improves plant growth and stress tolerance. Cells, 10(3), 582.
[CrossRef] [Google Scholar] - Kandoi, D., Mohanty, S., Govindjee, & Tripathy, B. C. (2016). Towards efficient photosynthesis: overexpression of Zea mays phosphoenolpyruvate carboxylase in Arabidopsis thaliana. Photosynthesis Research, 130(1), 47-72.
[CrossRef] [Google Scholar] - Komatsu, S., Nakamura, T., Sugimoto, Y., & Sakamoto, K. (2014). Proteomic and metabolomic analyses of soybean root tips under flooding stress. Protein and Peptide Letters, 21(9), 865-884.
[CrossRef] [Google Scholar] - Chiang, C. M., Chen, C. C., Chen, S. P., Lin, K. H., Chen, L. R., Su, Y. H., & Yen, H. C. (2017). Overexpression of the ascorbate peroxidase gene from eggplant and sponge gourd enhances flood tolerance in transgenic Arabidopsis. Journal of plant research, 130(2), 373-386.
[CrossRef] [Google Scholar] - Zhang, J., Yuan, H., Li, Y., Chen, Y., Liu, G., Ye, M., ... & Xu, J. (2020). Genome sequencing and phylogenetic analysis of allotetraploid Salix matsudana Koidz. Horticulture Research, 7(1), 201.
[CrossRef] [Google Scholar] - Chen, C., Chen, H., Zhang, Y., Thomas, H. R., Frank, M. H., He, Y., & Xia, R. (2020). TBtools: an integrative toolkit developed for interactive analyses of big biological data. Molecular Plant, 13(8), 1194-1202.
[CrossRef] [Google Scholar] - Huang, Q., Hua, X., Zhang, Q., Pan, W., Wang, Y., Liu, G., ... & Zhang, J. (2023). Identification and functional verification of salt tolerance hub genes in Salix matsudana based on QTL and transcriptome analysis. Environmental and Experimental Botany, 215, 105470.
[CrossRef] [Google Scholar] - Chen, Y., Yang, J., Guo, H., Du, Y., Liu, G., Yu, C., ... & Zhang, J. (2022). Comparative transcriptomic analysis reveals potential mechanisms for high tolerance to submergence in arbor willows. PeerJ, 10, e12881.
[CrossRef] [Google Scholar] - Izui, K., Matsumura, H., Furumoto, T., & Kai, Y. (2004). Phosphoenolpyruvate carboxylase: a new era of structural biology. Annual Review of Plant Biology, 55(1), 69-84.
[CrossRef] [Google Scholar] - O’Leary, B., Fedosejevs, E. T., Hill, A. T., Bettridge, J., Park, J., Rao, S. K., ... & Plaxton, W. C. (2011). Tissue-specific expression and post-translational modifications of plant-and bacterial-type phosphoenolpyruvate carboxylase isozymes of the castor oil plant, Ricinus communis L. Journal of Experimental Botany, 62(15), 5485-5495.
[CrossRef] [Google Scholar] - Ryan, P. R., Delhaize, E., & Jones, D. L. (2001). Function and mechanism of organic anion exudation from plant roots. Annual Review of Plant Biology, 52(1), 527-560.
[CrossRef] [Google Scholar] - Heyduk, K., Moreno-Villena, J. J., Gilman, I. S., Christin, P. A., & Edwards, E. J. (2019). The genetics of convergent evolution: insights from plant photosynthesis. Nature Reviews Genetics, 20(8), 485-493.
[CrossRef] [Google Scholar] - O'Leary, B., & Plaxton, W. C. (2020). Multifaceted functions of post-translational enzyme modifications in the control of plant glycolysis. Current Opinion in Plant Biology, 55, 28-37.
[CrossRef] [Google Scholar] - Wei, H., Lu, Z., Jiang, H., Xue, C., Xu, X., Liu, G., ... & Zhang, J. (2025). Comprehensive analysis of PEPC gene family in Populus trichocarpa: Characterization, evolutionary insights, and the role of PtPEPC4-PtLTPG14 interaction in carbon metabolism. Plant Physiology and Biochemistry, 220, 109573.
[CrossRef] [Google Scholar] - Sánchez, R., & Cejudo, F. J. (2003). Identification and expression analysis of a gene encoding a bacterial-type phosphoenolpyruvate carboxylase from Arabidopsis and rice. Plant Physiology, 132(2), 949-957.
[CrossRef] [Google Scholar] - Flagel, L. E., & Wendel, J. F. (2009). Gene duplication and evolutionary novelty in plants. New Phytologist, 183(3), 557-564.
[CrossRef] [Google Scholar] - Khan, F., Siddique, A. B., Shabala, S., Zhou, M., & Zhao, C. (2023). Phosphorus plays key roles in regulating plants’ physiological responses to abiotic stresses. Plants, 12(15), 2861.
[CrossRef] [Google Scholar] - Qiao, X., Zhang, S., & Paterson, A. H. (2022). Pervasive genome duplications across the plant tree of life and their links to major evolutionary innovations and transitions. Computational and structural biotechnology journal, 20, 3248-3256.
[CrossRef] [Google Scholar] - Wang, Y., Tan, X., & Paterson, A. H. (2013). Different patterns of gene structure divergence following gene duplication in Arabidopsis. BMC genomics, 14(1), 652.
[CrossRef] [Google Scholar] - Uhrig, R. G., O'Leary, B., Spang, H. E., MacDonald, J. A., She, Y. M., & Plaxton, W. C. (2008). Coimmunopurification of phosphorylated bacterial-and plant-type phospho enol pyruvate carboxylases with the plastidial pyruvate dehydrogenase complex from developing castor oil seeds. Plant Physiology, 146(3), 1346-1357.
[CrossRef] [Google Scholar] - Igawa, T., Fujiwara, M., Tanaka, I., Fukao, Y., & Yanagawa, Y. (2010). Characterization of bacterial-type phosphoenolpyruvate carboxylase expressed in male gametophyte of higher plants. BMC Plant Biology, 10(1), 200.
[CrossRef] [Google Scholar] - O'Leary, B., Rao, S. K., Kim, J., & Plaxton, W. C. (2009). Bacterial-type phosphoenolpyruvate carboxylase (PEPC) functions as a catalytic and regulatory subunit of the novel class-2 PEPC complex of vascular plants. Journal of Biological Chemistry, 284(37), 24797-24805.
[CrossRef] [Google Scholar] - Park, J., Khuu, N., Howard, A. S., Mullen, R. T., & Plaxton, W. C. (2012). Bacterial‐and plant‐type phosphoenolpyruvate carboxylase isozymes from developing castor oil seeds interact in vivo and associate with the surface of mitochondria. The Plant Journal, 71(2), 251-262.
[CrossRef] [Google Scholar] - Behera, D., Swain, A., Karmakar, S., Dash, M., Swain, P., Baig, M. J., & Molla, K. A. (2023). Overexpression of Setaria italica phosphoenolpyruvate carboxylase gene in rice positively impacts photosynthesis and agronomic traits. Plant Physiology and Biochemistry, 194, 169-181.
[CrossRef] [Google Scholar] - Sánchez, R., Flores, A., & Cejudo, F. J. (2006). Arabidopsis phosphoenolpyruvate carboxylase genes encode immunologically unrelated polypeptides and are differentially expressed in response to drought and salt stress. Planta, 223(5), 901-909.
[CrossRef] [Google Scholar]
Cite This Article
TY - JOUR AU - Wei, Hui AU - Cao, Yi AU - Zhou, Xiaoxi AU - Sun, Kaixin AU - Shan, Yuxin AU - Cao, Tongtong AU - Liu, Guoyuan AU - Lian, Bolin AU - Zhong, Fei AU - Shi, Jian AU - Zhang, Jian PY - 2026 DA - 2026/03/17 TI - Characterization, Evolutionary Insights, and Stress-Responsive Expression of the Phosphoenolpyruvate Carboxylase (PEPC) Gene Family in Salix Matsudana JO - Plant Innovation Journal T2 - Plant Innovation Journal JF - Plant Innovation Journal VL - 1 IS - 1 SP - 27 EP - 49 DO - 10.62762/PIJ.2025.111778 UR - https://www.icck.org/article/abs/PIJ.2025.111778 KW - Salix matsudana KW - PEPC KW - synteny analysis KW - salt or submergence stress AB - Phosphoenolpyruvate carboxylase (PEPC) is a vital enzyme in plant carbon metabolism, catalyzing the conversion of phosphoenolpyruvate (PEP) to oxaloacetate (OAA), a key intermediate in numerous biosynthetic pathways. PEPC family members are usually classified into PTPC and BTPC subfamilies, and PTPC subfamily is characterized by a conserved N-terminal serine phosphorylation site, while BTPC lacks this site. In this study, we identified 10 Salix matsudana PEPC genes (SmPEPCs), which were classified into two subfamilies, PTPC and BTPC, based on phylogenetic tree. Our findings highlighted significant gene duplication events that contributed to the expansion of the SmPEPC gene family, shedding light on its evolutionary development in willow. Comparative synteny analysis with Arabidopsis thaliana, Oryza sativa, Populus trichocarpa, and S. purpurea revealed both conserved and diverged patterns of PEPC organization across species. Interaction network analysis, along with Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis, indicated that SmPEPCs are involved in crucial metabolic pathways related to carbon flux. Expression profiling under salt or submergence stress revealed distinct expression patterns of SmPEPCs. Specifically, SmPEPC9 showed predominant upregulation under salt stress, while SmPEPC6 was mainly upregulated in submergence-tolerant varieties under submergence stress. These results suggested that SmPEPCs potentially modulate carbon flux and organic acid metabolism to mitigate stress effects, playing a central role in stress adaptation mechanisms. SN - pending PB - Institute of Central Computation and Knowledge LA - English ER -
@article{Wei2026Characteri,
author = {Hui Wei and Yi Cao and Xiaoxi Zhou and Kaixin Sun and Yuxin Shan and Tongtong Cao and Guoyuan Liu and Bolin Lian and Fei Zhong and Jian Shi and Jian Zhang},
title = {Characterization, Evolutionary Insights, and Stress-Responsive Expression of the Phosphoenolpyruvate Carboxylase (PEPC) Gene Family in Salix Matsudana},
journal = {Plant Innovation Journal},
year = {2026},
volume = {1},
number = {1},
pages = {27-49},
doi = {10.62762/PIJ.2025.111778},
url = {https://www.icck.org/article/abs/PIJ.2025.111778},
abstract = {Phosphoenolpyruvate carboxylase (PEPC) is a vital enzyme in plant carbon metabolism, catalyzing the conversion of phosphoenolpyruvate (PEP) to oxaloacetate (OAA), a key intermediate in numerous biosynthetic pathways. PEPC family members are usually classified into PTPC and BTPC subfamilies, and PTPC subfamily is characterized by a conserved N-terminal serine phosphorylation site, while BTPC lacks this site. In this study, we identified 10 Salix matsudana PEPC genes (SmPEPCs), which were classified into two subfamilies, PTPC and BTPC, based on phylogenetic tree. Our findings highlighted significant gene duplication events that contributed to the expansion of the SmPEPC gene family, shedding light on its evolutionary development in willow. Comparative synteny analysis with Arabidopsis thaliana, Oryza sativa, Populus trichocarpa, and S. purpurea revealed both conserved and diverged patterns of PEPC organization across species. Interaction network analysis, along with Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis, indicated that SmPEPCs are involved in crucial metabolic pathways related to carbon flux. Expression profiling under salt or submergence stress revealed distinct expression patterns of SmPEPCs. Specifically, SmPEPC9 showed predominant upregulation under salt stress, while SmPEPC6 was mainly upregulated in submergence-tolerant varieties under submergence stress. These results suggested that SmPEPCs potentially modulate carbon flux and organic acid metabolism to mitigate stress effects, playing a central role in stress adaptation mechanisms.},
keywords = {Salix matsudana, PEPC, synteny analysis, salt or submergence stress},
issn = {pending},
publisher = {Institute of Central Computation and Knowledge}
}
Article Metrics
Publisher's Note
ICCK stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and Permissions
 Copyright © 2026 by the Author(s). Published by Institute of Central Computation and Knowledge. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.
Copyright © 2026 by the Author(s). Published by Institute of Central Computation and Knowledge. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.


