Abstract
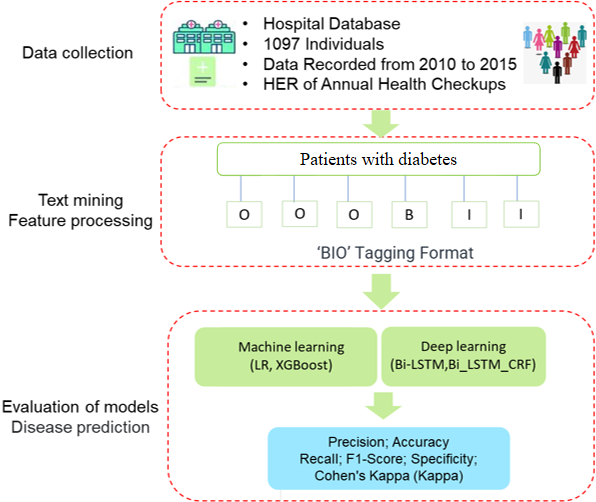
In the healthcare sector, the application of deep learning technologies has revolutionized data analysis and disease forecasting. This is particularly evident in the field of diabetes, where the deep analysis of Electronic Health Records (EHR) has unlocked new opportunities for early detection and effective intervention strategies. Our research presents an innovative model that synergizes the capabilities of Bidirectional Long Short-Term Memory Networks-Conditional Random Field (BiLSTM-CRF) with a fusion of XGBoost and Logistic Regression. This model is designed to enhance the accuracy of diabetes risk prediction by conducting an in-depth analysis of electronic medical records data. The first phase of our approach involves employing BiLSTM-CRF to delve into the temporal characteristics and latent patterns present in EHR data. This method effectively uncovers the progression trends of diabetes, which are often hidden in the complex data structures of medical records. The second phase leverages the combined strength of XGBoost and Logistic Regression to classify these extracted features and evaluate associated risks. This dual approach facilitates a more nuanced and precise prediction of diabetes, outperforming traditional models, particularly in handling multifaceted and nonlinear medical datasets. Our research demonstrates a notable advancement in diabetes prediction over traditional methods, showcasing the effectiveness of our combined BiLSTM-CRF, XGBoost, and Logistic Regression model. This study highlights the value of data-driven strategies in clinical decision-making, equipping healthcare professionals with precise tools for early detection and intervention. By enabling personalized treatment and timely care, our approach signifies progress in incorporating advanced analytics in healthcare, potentially improving outcomes for diabetes and other chronic conditions.
Keywords
deep learning
electronic health records
BiLSTM-CRF
XGBoost
healthcare analytics
Funding
This work was supported without any funding.
Cite This Article
APA Style
Pang, H., Zhou, L., Dong, Y., Chen, P., Gu, D., Lyu, T. & Zhang, H. (2024). Electronic Health Records-Based Data-Driven Diabetes Knowledge Unveiling and Risk Prognosis. ICCK Transactions on Intelligent Systematics, 2(1), 1–13. https://doi.org/10.62762/TIS.2025.367320
Publisher's Note
ICCK stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Institute of Central Computation and Knowledge (ICCK) or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.


 Submit Manuscript
Edit a Special Issue
Submit Manuscript
Edit a Special Issue