Rational Design of Plant Chemical Factories: CRISPR-Based Metabolic Engineering in Medicinal and Aromatic Plants

Article Information
Abstract
Medicinal and aromatic plants (MAPs) serve as biochemical factories producing valuable secondary metabolites, yet their potential is limited by low yields, tissue-specific accumulation, and co-production of toxic compounds. Traditional improvement methods have achieved only incremental gains, highlighting the need for precision metabolic engineering. CRISPR/Cas genome editing has revolutionized this field by enabling targeted modifications from gene knockouts to single-nucleotide changes. This review examines core strategies in applying genome editing to engineer MAP metabolic pathways, including gene disruption, transcriptional modulation, and multiplex editing to redirect flux, eliminate competing pathways, and remove toxic branches. Case studies demonstrate successes in alkaloid engineering—such as clean chemotypes with pure hyoscyamine in Atropa belladonna (>50% yield increase) and detoxified Symphytum officinale—and terpenoid enhancement with three-fold glycyrrhizin increase in licorice through combined blocking and overexpression. These examples showcase CRISPR's surgical precision for improving pharmaceutical purity, safety, and production efficiency. We discuss integration with multi-omics for systems-level optimization and explore emerging frontiers including base editing, prime editing, and CRISPR-directed evolution for enzyme optimization. As these technologies mature, genome editing will transform MAPs into customizable cellular factories delivering sustainable, high-value bioproducts.
Graphical Abstract
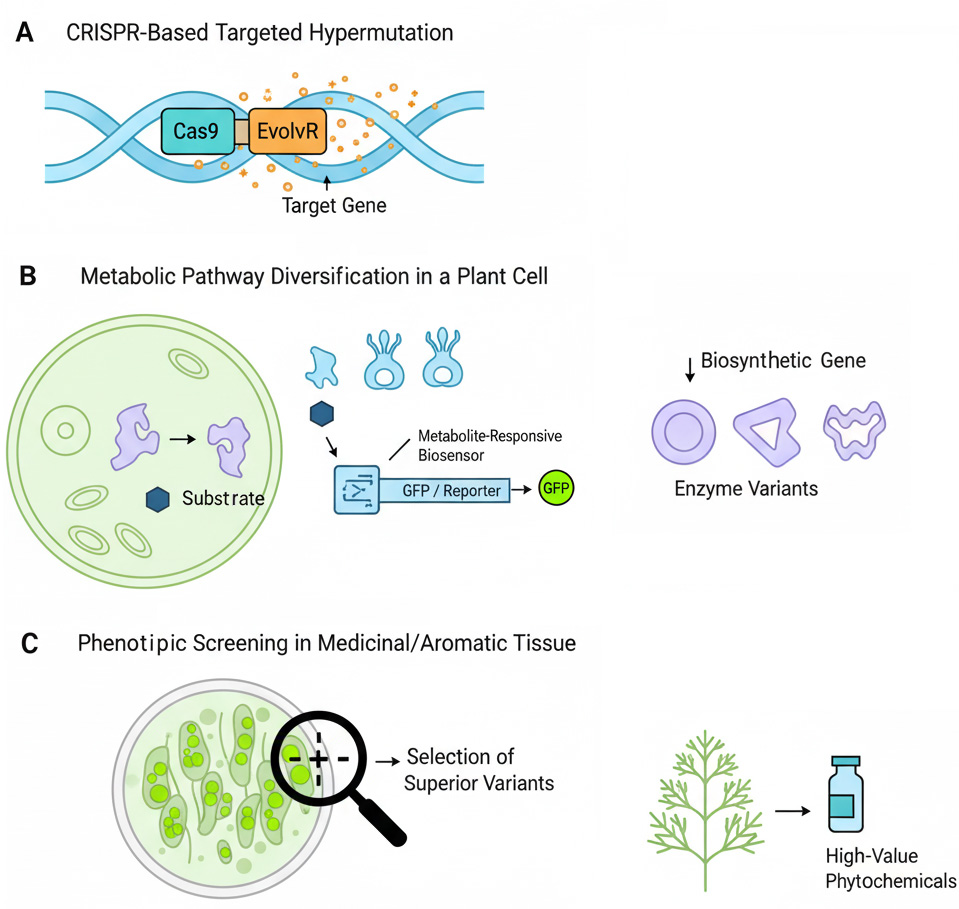
Keywords
Data Availability Statement
Funding
Conflicts of Interest
AI Use Statement
Ethical Approval and Consent to Participate
References
- Borah, A., Singh, S., Chattopadhyay, R., Kaur, J., & Bari, V. K. (2024). Integration of CRISPR/Cas9 with multi-omics technologies to engineer secondary metabolite productions in medicinal plant: Challenges and Prospects. Functional & Integrative Genomics, 24(6), 207.
[CrossRef] [Google Scholar] - Shabir, P. A. (2021). CRISPR/Cas9-mediated genome editing in medicinal and aromatic plants: developments and applications. In Medicinal and aromatic plants (pp. 209-221). Academic Press.
[CrossRef] [Google Scholar] - Shelepova, O. V., Baranova, E. N., Tkacheva, E. V., Evdokimenkova, Y. B., Ivanovskii, A. A., Konovalova, L. N., & Gulevich, A. A. (2022). Aromatic plants metabolic engineering: a review. Agronomy, 12(12), 3131.
[CrossRef] [Google Scholar] - Naik, B. J., Vasamsetti, B. M. K., Kim, S. C., Gunti, M., Swamy, M. K., & Mekapogu, M. (2023). Genome editing in medicinal plants using CRISPR/Cas9 tool. In Phytochemical genomics: plant metabolomics and medicinal plant genomics (pp. 571-589). Singapore: Springer Nature Singapore.
[CrossRef] [Google Scholar] - Zeng, L., Zhang, Q., Jiang, C., Zheng, Y., Zuo, Y., Qin, J., ... & Deng, H. (2021). Development of Atropa belladonna L. plants with high-yield hyoscyamine and without its derivatives using the CRISPR/Cas9 system. International Journal of Molecular Sciences, 22(4), 1731.
[CrossRef] [Google Scholar] - Zakaria, M. M., Schemmerling, B., & Ober, D. (2021). CRISPR/Cas9-mediated genome editing in comfrey (Symphytum officinale) hairy roots results in the complete eradication of pyrrolizidine alkaloids. Molecules, 26(6), 1498.
[CrossRef] [Google Scholar] - Kandoudi, W., & Németh-Zámboriné, É. (2022). Stimulating secondary compound accumulation by elicitation: Is it a realistic tool in medicinal plants in vivo?. Phytochemistry Reviews, 21(6), 2007-2025.
[CrossRef] [Google Scholar] - Zhu, S., Fu, R., Martin, C., Zhao, Q., & Zhang, Y. (2025). Recent advances in biosynthesis, regulation, and metabolic engineering of flavonoids. Plant Physiology, 199(1), kiaf356.
[CrossRef] [Google Scholar] - Sabzehzari, M., Zeinali, M., & Naghavi, M. R. (2020). CRISPR-based metabolic editing: Next-generation metabolic engineering in plants. Gene, 759, 144993.
[CrossRef] [Google Scholar] - Cao, Y., Zhou, H., Zhou, X., & Li, F. (2020). Control of plant viruses by CRISPR/Cas system-mediated adaptive immunity. Frontiers in Microbiology, 11, 593700.
[CrossRef] [Google Scholar] - Wang, Y., Zafar, N., Ali, Q., Manghwar, H., Wang, G., Yu, L., ... & Jin, S. (2022). CRISPR/Cas genome editing technologies for plant improvement against biotic and abiotic stresses: advances, limitations, and future perspectives. Cells, 11(23), 3928.
[CrossRef] [Google Scholar] - Čermák, T., Curtin, S. J., Gil-Humanes, J., Čegan, R., Kono, T. J., Konečná, E., ... & Voytas, D. F. (2017). A multipurpose toolkit to enable advanced genome engineering in plants. The plant cell, 29(6), 1196-1217.
[CrossRef] [Google Scholar] - Adli, M. (2018). The CRISPR tool kit for genome editing and beyond. Nature Communications, 9(1), 1911.
[CrossRef] [Google Scholar] - Chiyo, N., Seki, H., Kanamoto, T., Ueda, H., Kojoma, M., & Muranaka, T. (2024). Glycyrrhizin Production in Licorice Hairy Roots Based on Metabolic Redirection of Triterpenoid Biosynthetic Pathway by Genome Editing. Plant and Cell Physiology, 65(2), 185–198.
[CrossRef] [Google Scholar] - Pireyre, M., & Burow, M. (2015). Regulation of MYB and bHLH transcription factors: a glance at the protein level. Molecular Plant, 8(3), 378–388.
[CrossRef] [Google Scholar] - Zhou, Y., Feng, J., Li, Q., Huang, D., Chen, X., Du, Z., ... & Chen, W. (2020). Sm MYC2b enhances tanshinone accumulation in Salvia miltiorrhiza by activating pathway genes and promoting lateral root development. Frontiers in Plant Science, 11, 559438.
[CrossRef] [Google Scholar] - Liu, S., Wang, Y., Shi, M., Maoz, I., Gao, X., Sun, M., ... & Kai, G. (2022). SmbHLH60 and SmMYC2 antagonistically regulate phenolic acids and anthocyanins biosynthesis in Salvia miltiorrhiza. Journal of Advanced Research, 42, 205-219.
[CrossRef] [Google Scholar] - Han, C., Dong, X., Xing, X., Wang, Y., Feng, X., Sang, W., ... & He, Y. (2024). Gibberellin-Induced Transcription Factor Sm MYB71 Negatively Regulates Salvianolic Acid Biosynthesis in Salvia miltiorrhiza. Molecules, 29(24), 5892.
[CrossRef] [Google Scholar] - Shen, Q., Huang, H., Xie, L., Hao, X., Kayani, S. I., Liu, H., ... & Tang, K. (2022). Basic helix-loop-helix transcription factors AabHLH2 and AabHLH3 function antagonistically with AaMYC2 and are negative regulators in artemisinin biosynthesis. Frontiers in Plant Science, 13, 885622.
[CrossRef] [Google Scholar] - Nakayasu, M., Akiyama, R., Lee, H. J., Osakabe, K., Osakabe, Y., Watanabe, B., ... & Mizutani, M. (2018). Generation of $\alpha$-solanine-free hairy roots of potato by CRISPR/Cas9 mediated genome editing of the St16DOX gene. Plant Physiology and Biochemistry, 131, 70-77.
[CrossRef] [Google Scholar] - Hasebe, F., Yuba, H., Hashimoto, T., Saito, K., Funa, N., & Shoji, T. (2021). CRISPR/Cas9-mediated disruption of the PYRROLIDINE KETIDE SYNTHASE gene reduces the accumulation of tropane alkaloids in Atropa belladonna hairy roots. Bioscience, biotechnology, and biochemistry, 85(12), 2404-2409.
[CrossRef] [Google Scholar] - Anzalone, A. V., Randolph, P. B., Davis, J. R., Sousa, A. A., Koblan, L. W., Levy, J. M., ... & Liu, D. R. (2019). Search-and-replace genome editing without double-strand breaks or donor DNA. Nature, 576(7785), 149-157.
[CrossRef] [Google Scholar] - Thimmappa, R., Geisler, K., Louveau, T., O'Maille, P., & Osbourn, A. (2014). Triterpene biosynthesis in plants. Annual Review of Plant Biology, 65, 225–257.
[CrossRef] [Google Scholar] - Liu, Y., Zhou, J., Hu, T., Lu, Y., Gao, L., Tu, L., & others. (2020). Identification and functional characterization of squalene epoxidases and oxidosqualene cyclases from Tripterygium wilfordii. Plant Cell Reports, 39(3), 409–418.
[CrossRef] [Google Scholar] - Halperin, S. O., Tou, C. J., Wong, E. B., Modavi, C., Schaffer, D. V., & Dueber, J. E. (2018). CRISPR-guided DNA polymerases enable diversification of all nucleotides in a tunable window. Nature, 560(7717), 248–252.
[CrossRef] [Google Scholar] - d’Oelsnitz, S., Kim, W., Burkholder, N. T., Javanmardi, K., Thyer, R., Zhang, Y., ... & Ellington, A. D. (2022). Using fungible biosensors to evolve improved alkaloid biosyntheses. Nature chemical biology, 18(9), 981-989.
[CrossRef] [Google Scholar]
Cite This Article
TY - JOUR AU - Kadkhodaei, Saeid AU - Shahabadi, Hassan Zadabbas AU - Hosseini-Monfared, Hossein PY - 2026 DA - 2026/03/12 TI - Rational Design of Plant Chemical Factories: CRISPR-Based Metabolic Engineering in Medicinal and Aromatic Plants JO - Plant Innovation Journal T2 - Plant Innovation Journal JF - Plant Innovation Journal VL - 1 IS - 1 SP - 18 EP - 26 DO - 10.62762/PIJ.2025.283212 UR - https://www.icck.org/article/abs/PIJ.2025.283212 KW - CRISPR/Cas9 KW - metabolic engineering KW - medicinal plants KW - alkaloid biosynthesis KW - terpenoid biosynthesis KW - gene knockout KW - pathway optimization KW - secondary metabolism AB - Medicinal and aromatic plants (MAPs) serve as biochemical factories producing valuable secondary metabolites, yet their potential is limited by low yields, tissue-specific accumulation, and co-production of toxic compounds. Traditional improvement methods have achieved only incremental gains, highlighting the need for precision metabolic engineering. CRISPR/Cas genome editing has revolutionized this field by enabling targeted modifications from gene knockouts to single-nucleotide changes. This review examines core strategies in applying genome editing to engineer MAP metabolic pathways, including gene disruption, transcriptional modulation, and multiplex editing to redirect flux, eliminate competing pathways, and remove toxic branches. Case studies demonstrate successes in alkaloid engineering—such as clean chemotypes with pure hyoscyamine in Atropa belladonna (>50% yield increase) and detoxified Symphytum officinale—and terpenoid enhancement with three-fold glycyrrhizin increase in licorice through combined blocking and overexpression. These examples showcase CRISPR's surgical precision for improving pharmaceutical purity, safety, and production efficiency. We discuss integration with multi-omics for systems-level optimization and explore emerging frontiers including base editing, prime editing, and CRISPR-directed evolution for enzyme optimization. As these technologies mature, genome editing will transform MAPs into customizable cellular factories delivering sustainable, high-value bioproducts. SN - pending PB - Institute of Central Computation and Knowledge LA - English ER -
@article{Kadkhodaei2026Rational,
author = {Saeid Kadkhodaei and Hassan Zadabbas Shahabadi and Hossein Hosseini-Monfared},
title = {Rational Design of Plant Chemical Factories: CRISPR-Based Metabolic Engineering in Medicinal and Aromatic Plants},
journal = {Plant Innovation Journal},
year = {2026},
volume = {1},
number = {1},
pages = {18-26},
doi = {10.62762/PIJ.2025.283212},
url = {https://www.icck.org/article/abs/PIJ.2025.283212},
abstract = {Medicinal and aromatic plants (MAPs) serve as biochemical factories producing valuable secondary metabolites, yet their potential is limited by low yields, tissue-specific accumulation, and co-production of toxic compounds. Traditional improvement methods have achieved only incremental gains, highlighting the need for precision metabolic engineering. CRISPR/Cas genome editing has revolutionized this field by enabling targeted modifications from gene knockouts to single-nucleotide changes. This review examines core strategies in applying genome editing to engineer MAP metabolic pathways, including gene disruption, transcriptional modulation, and multiplex editing to redirect flux, eliminate competing pathways, and remove toxic branches. Case studies demonstrate successes in alkaloid engineering—such as clean chemotypes with pure hyoscyamine in Atropa belladonna (>50\% yield increase) and detoxified Symphytum officinale—and terpenoid enhancement with three-fold glycyrrhizin increase in licorice through combined blocking and overexpression. These examples showcase CRISPR's surgical precision for improving pharmaceutical purity, safety, and production efficiency. We discuss integration with multi-omics for systems-level optimization and explore emerging frontiers including base editing, prime editing, and CRISPR-directed evolution for enzyme optimization. As these technologies mature, genome editing will transform MAPs into customizable cellular factories delivering sustainable, high-value bioproducts.},
keywords = {CRISPR/Cas9, metabolic engineering, medicinal plants, alkaloid biosynthesis, terpenoid biosynthesis, gene knockout, pathway optimization, secondary metabolism},
issn = {pending},
publisher = {Institute of Central Computation and Knowledge}
}
Article Metrics
Publisher's Note
ICCK stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and Permissions
 Copyright © 2026 by the Author(s). Published by Institute of Central Computation and Knowledge. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.
Copyright © 2026 by the Author(s). Published by Institute of Central Computation and Knowledge. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.


